基于CVAE的二维分子条件生成
上篇文章我们实现了基于VAE架构的分子生成,他支持有上限原子的分子随机生成,接下来我们将学习如何引入条件控制,从而做到生成给定条件(分子性质)范围的分子,本文项目结构如下:
CVAE 2D/
├── data/ # 数据集文件
│ └── ZINC.csv
├── result/ # 结果目录
│ ├── model.pth # 模型权重
│ ├── result.csv # 推理生成的结果
│ └── losses.csv # 平均损失记录
├── config.py # 超参数配置文件
├── dataset.py # 加载数据集
├── model.py # 训练代码
├── generate.py # 生成代码
└── draw.py # 绘图分析脚本项目结构和VAE版本大致相同,我们把训练和推理的代码分开实现,同时补充进行数据分析的脚本函数。
本文基于该文章进行了代码优化和改造:
参考文献:MGCVAE: Multi-Objective Inverse Design via Molecular Graph Conditional Variational Autoencoder,文章链接在此处
配置信息
以下文件在config.py中实现,这些和VAE生成的配置完全一致。
import torch
import torch.nn as nn
from rdkit import Chem
from rdkit.Chem import rdchem
from pathlib import Path
result_path = Path('./result')
result_path.mkdir(exist_ok=True, parents=True)
# 数据集路径
DATA_FILE = "./data/ZINC.csv"
# 权重保存文件
PT_PATH = "./result/model.pth"
# 损失保存路径
LOSS_PATH = "./result/losses.csv"
# 推理生成结果文件
RESTULT_PATH = "./result/result"
# 测试集/训练集划分比例
DIVIDE_RATIO = 0.1主要增加了CONDITIONS字段,用于配置需要生成的“条件”约束,这里我们约束的是logP和MR值。
# 要生成的分子数
GEN_NUM = 10000
# 生成条件
CONDITIONS = {
'logP_list':[0, -1, -2, -3],
'mr_list': [2, 3, 4, 5, 6]
}模型配置和超参数等也和之前的完全一致,最后增加了COND_DIM条件编码向量的维度,这里设置为16。
# 批次大小
BATCH_SIZE = 1000
# 训练周期
EPOCH = 200
# 学习率
LR = 0.00005
# 设备
DEVICE = "cuda" if torch.cuda.is_available() else "cpu"
# 支持的原子类型列表
ATOM_LIST = ( 'C', 'O', 'N', 'H', 'P', 'S', 'F', 'Cl', 'Br', 'I', 'B', 'Si', 'Sn')
ATOM_LIST = [0] + [Chem.Atom(x).GetAtomicNum() for x in ATOM_LIST]
# 支持的原子种类数
ATOM_LEN = len(ATOM_LIST)
# 支持的键类型列表
BOND_LIST = (
rdchem.BondType.ZERO,
rdchem.BondType.SINGLE,
rdchem.BondType.DOUBLE,
rdchem.BondType.TRIPLE,
rdchem.BondType.AROMATIC
)
# 支持的键类型数
BOND_LEN = len(BOND_LIST)
# 索引和实际类型对照表
atom_encoder_m = {l: i for i, l in enumerate(ATOM_LIST)}
atom_decoder_m = {i: l for i, l in enumerate(ATOM_LIST)}
bond_encoder_m = {l: i for i, l in enumerate(BOND_LIST)}
bond_decoder_m = {i: l for i, l in enumerate(BOND_LIST)}
# VAE参数配置
# Encoder: IN_DIM -> H1_DIM -> H2_DIM -> Z_DIM
# Decoder: Z_DIM -> H2_DIM -> H1_DIM -> IN_DIM (out)
# 最大原子数
MAX_SIZE = 16
# 分子特征矩阵列数
F_COL = 1 + ATOM_LEN + MAX_SIZE * BOND_LEN
# 输入维度
IN_DIM = MAX_SIZE * F_COL
# 隐空间维度
Z_DIM = 128
# 隐藏层1
H1_DIM = 512
# 隐藏层2
H2_DIM = 256
# 条件编码维度
COND_DIM = 16数据集加载
该部分代码在dataset.py中实现。
节点特征矩阵、边特征矩阵和mol对象转特征矩阵的函数和VAE部分完全一致。
import pandas as pd
from rdkit import Chem
import torch
from torch.utils.data import Dataset, DataLoader
from config import *
import numpy as np
# 生成节点特征矩阵
def node_features(mol, max_length=MAX_SIZE, atom_types=ATOM_LEN):
# 预制全零张量
features = np.zeros([max_length, atom_types], dtype=np.float32)
# 填充真实原子
for i, atom in enumerate(mol.GetAtoms()):
atomic_num = atom.GetAtomicNum()
if atomic_num in atom_encoder_m:
features[i, atom_encoder_m[atomic_num]] = 1.0
# 多出的原子标记为0号(虚拟原子)
actual_atoms = min(mol.GetNumAtoms(), max_length)
features[actual_atoms:, 0] = 1.0
return torch.tensor(features, dtype=torch.float32)
# 生成边特征矩阵
def bond_features(mol, bond_encoder_m, max_length=MAX_SIZE):
bond_types = len(bond_encoder_m)
# 预先构建三维张量,所有键均为“非键”编码的独热向量
A = torch.zeros(bond_types, dtype=torch.int32)
A[0] = 1
A = A.expand(max_length, max_length, bond_types).clone()
# 遍历每个边
for bond in mol.GetBonds():
begin = bond.GetBeginAtomIdx() # 键的起始原子索引
end = bond.GetEndAtomIdx() # 键的终止原子索引
i = min(begin, end) # 行号=原子对中较小的索引
j = max(begin, end) # 列号=原子对中较大的索引
# 构建边类型独热编码向量,替换原值
bond_code = bond_encoder_m[bond.GetBondType()]
bond_onehot = torch.zeros(bond_types, dtype=torch.int32)
bond_onehot[bond_code] = 1
A[i, j] = bond_onehot
# 将三维矩阵按最后一维展开,得到二维特征矩阵
A = A.reshape(max_length, -1)
return A
# mol对象生成分子特征矩阵
def mol_to_graph(mol, length, smiles='', max_num=MAX_SIZE):
try:
mol_length = int(length) - 1
vec_length = torch.zeros([max_num,1], dtype=torch.float)
vec_length[mol_length, 0] = 1.0
graph = torch.cat([vec_length, node_features(mol, max_num),bond_features(mol, bond_encoder_m)], 1).float()
return graph
except Exception as e:
print('Error:', smiles, 'more information:', e)随后是读取CSV数据集的内容,创建mol对象,和CVAE也完全一致。
# 读取数据集源文件
df = pd.read_csv(DATA_FILE)
# 按设定大小抽样
df = df[df['Length'] <= MAX_SIZE].reset_index(drop=True)
# 生成mol对象列
df["mol"] = df["SMILES"].apply(lambda x: Chem.MolFromSmiles(x))
# 去除空值
df = df[df["mol"].notna()].reset_index(drop=True)
# 生成特征矩阵列
df['graph'] = df[['mol', 'Length', 'SMILES']].apply(
lambda row: mol_to_graph(mol=row['mol'], length=row['Length'], smiles=row['SMILES']),
axis=1
)
# 去除空值
df = df[df['graph'].notna()].reset_index(drop=True)不同的是,这里我们把df对象传入数据集类,构建get、len方法,不同的是这里返回的是特征矩阵、logP、MR三个张量。
# 定义数据集类
class MolGraphDataset(Dataset):
def __init__(self, df):
self.df = df.reset_index(drop=True)
def __len__(self):
return self.df.shape[0]
def __getitem__(self, idx):
row = self.df.iloc[idx]
feat_tensor = row['graph']
logp_tensor = torch.tensor([row['logP']], dtype=torch.float32)
mr_tensor = torch.tensor([row['MR']], dtype=torch.float32)
return feat_tensor, logp_tensor, mr_tensor 划分数据集的内容也和VAE完全一致。
# 初始化自定义数据集
mol_dataset = MolGraphDataset(df)
# 划分训练/测试集
tr = 1 - DIVIDE_RATIO
train_size = int(len(mol_dataset) * tr)
test_size = len(mol_dataset) - train_size
# 随机划分
train_dataset, test_dataset = torch.utils.data.random_split(
mol_dataset,
[train_size, test_size],
generator=torch.Generator().manual_seed(42)
)
# 构建DataLoader
train_loader = DataLoader(
dataset=train_dataset,
batch_size=BATCH_SIZE,
shuffle=True,
drop_last=True,
)
test_loader = DataLoader(
dataset=test_dataset,
batch_size=BATCH_SIZE,
shuffle=False,
drop_last=True,
)对Dataloader进行单元测试,单批次会返回三个张量,看看他们的形状。
if __name__ == "__main__":
print(f"Train dataset size: {len(train_dataset)} samples")
print(f"Test dataset size: {len(test_dataset)} samples")
print(f"Train loader batch count: {len(train_loader)}")
print(f"Test loader batch count: {len(test_loader)}")
# 测试加载单个批次
for feat_tensor, logp_tensor, mr_tensor in train_loader:
print(f"Feature shape: {feat_tensor.shape}")
print(f"LogP shape: {logp_tensor.shape}")
print(f"MR shape: {mr_tensor.shape}")
break可以看到,全部10w条样本以9:1被划分为训练集和测试集,每批次1000个样本,特征矩阵
Train dataset size: 90000 samples
Test dataset size: 10000 samples
Train loader batch count: 90
Test loader batch count: 10
Feature shape: torch.Size([1000, 16, 95])
LogP shape: torch.Size([1000, 1])
MR shape: torch.Size([1000, 1])此外,我们可以查看数据集的分布情况:
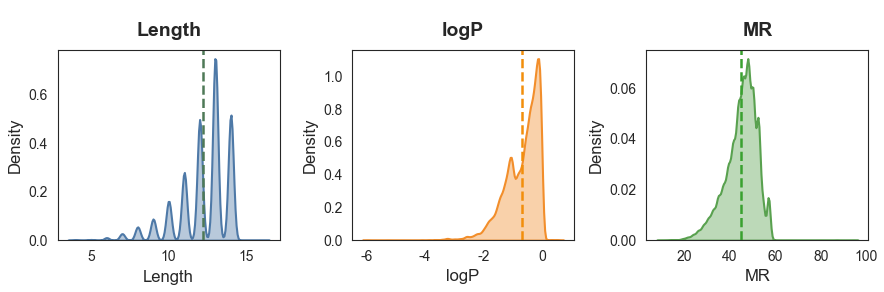
========== 各属性关键统计信息 ==========
Length logP MR
mean 12.20 -0.69 44.79
std 1.66 0.59 6.94
min 4.00 -5.97 9.70
max 16.00 0.58 94.42我们的数据集分子大小最大是16个原子,logP集中在[-6,1],MR集中在[20,60],因此配置MAX_SIZE = 16,COND_DIM = 16完全足够。
在生成时,条件限制在
CONDITIONS = {
'logP_list':[0, -1, -2, -3],
'mr_list': [2, 3, 4, 5, 6]
}模型定义
该部分代码在model.py中实现,完成模型的定义、损失函数计算和必要的工具函数。
损失函数依然使用KL散度+重构BCE损失,主要增加了生成条件编码向量的函数。
因为我们的属性值是连续特征,实际上这里是将其做了风箱划分,利用round四舍五入把连续特征变成离散特征,这样就可以使用独热编码向量来表征属性条件。
import torch
import torch.nn as nn
import torch.nn.functional as F
from config import *
# 损失函数
def loss_function(recon_x, x, mu, log_var):
# BCE损失
BCE = F.binary_cross_entropy(recon_x, x.view(-1, IN_DIM), reduction='sum')
# KL散度损失
KLD = -0.5 * torch.sum(1 + log_var - mu.pow(2) - log_var.exp())
return BCE + KLD
# 条件编码
def one_hot(labels, class_size=COND_DIM):
targets = torch.zeros(labels.shape[0], class_size, device=DEVICE)
for i, label in enumerate(labels):
targets[i, round(label.item())] = 1
return targets随后是模型的结构定义,因为我们是将分子图展平为一维向量送入VAE中,因此在有条件的情况下,只需要把多个条件向量拼接到分子图向量后即可,这样模型的输入维度将变成
# 模型定义
class CVAE(nn.Module):
def __init__(self, x_dim=IN_DIM, h_dim1=H1_DIM, h_dim2=H2_DIM, z_dim=Z_DIM, c_dim=COND_DIM):
super(CVAE, self).__init__()
self.x_dim = x_dim
# 编码器
self.fc1 = nn.Linear(x_dim+c_dim*2, h_dim1)
self.fc2 = nn.Linear(h_dim1, h_dim2)
self.fc31 = nn.Linear(h_dim2, z_dim)
self.fc32 = nn.Linear(h_dim2, z_dim)
# 解码器
self.fc4 = nn.Linear(c_dim*2+z_dim, h_dim2)
self.fc5 = nn.Linear(h_dim2, h_dim1)
self.fc6 = nn.Linear(h_dim1, x_dim)
# 在编码器的输入中将特征和条件拼接
def encoder(self, x, c1, c2):
concat_input = torch.cat([x, c1, c2], 1)
h = F.relu(self.fc1(concat_input))
h = F.relu(self.fc2(h))
return self.fc31(h), self.fc32(h)
def sampling(self, mu, log_var):
std = torch.exp(0.5*log_var)
eps = torch.randn_like(std)
return eps.mul(std).add(mu)
# 同样在解码器的输入中将特征和条件拼接
def decoder(self, z, c1, c2):
concat_input = torch.cat([z, c1, c2], 1)
h = F.relu(self.fc4(concat_input))
h = F.relu(self.fc5(h))
return torch.sigmoid(self.fc6(h))
def forward(self, x, c1, c2):
mu, log_var = self.encoder(x.view(-1, self.x_dim), c1, c2)
z = self.sampling(mu, log_var)
return self.decoder(z, c1, c2), mu, log_var训练
以下代码在train.py中实现,主要是在训练和测试函数中处理了属性值,对于mr值要缩放10倍,让其分布在[0,16]之间(不超出独热向量的范围)。
import pandas as pd
import torch.optim as optim
from config import *
from model import *
# 训练函数
def train(model, train_loader):
train_loss = 0
model.train()
for (graph, logp, mr) in train_loader:
graph = graph.to(DEVICE)
# 生成条件对应的one-hot向量,mr值要缩小10倍
logp = one_hot(logp).to(DEVICE)
mr = one_hot(mr/10).to(DEVICE)
optimizer.zero_grad()
recon_batch, mu, log_var = model(graph, logp, mr)
loss = loss_function(recon_batch, graph, mu, log_var)
loss.backward()
optimizer.step()
train_loss += loss.item()
return train_loss
# 测试函数
def test(model, test_loader):
test_loss= 0
model.eval()
with torch.no_grad():
for (graph, logp, mr) in test_loader:
graph = graph.to(DEVICE)
# 生成条件对应的one-hot向量
logp = one_hot(logp).to(DEVICE)
mr = one_hot(mr/10).to(DEVICE)
recon, mu, log_var = model(graph, logp, mr)
test_loss += loss_function(recon, graph, mu, log_var).item()
return test_loss开始训练模型:
if __name__ == "__main__":
# 定义模型,迁移设备
cvae = CVAE()
cvae.to(DEVICE)
# 设置优化器
optimizer = optim.Adam(cvae.parameters(), lr=LR)
print(f'Train device:{DEVICE}')
print('Load Dataset......')
# 载入数据集
from dataset import train_loader, test_loader
print('='*50)
print(f"Train dataset size: {len(train_loader.dataset)} samples")
print(f"Test dataset size: {len(test_loader.dataset)} samples")
print(f"Train loader batch count: {len(train_loader)}")
print(f"Test loader batch count: {len(test_loader)}")
print('='*50)
print('Training the model...')
# 开始迭代训练周期
train_loss_list = []
test_loss_list = []
for epoch in range(1, EPOCH+1):
# 训练周期损失
train_loss = train(cvae, train_loader) / len(train_loader.dataset)
train_loss_list.append(train_loss)
# 测试周期损失
test_loss = test(cvae, test_loader) / len(test_loader.dataset)
test_loss_list.append(test_loss)
print(f'Epoch [{epoch}/{EPOCH}], train loss {train_loss:.4f}; val loss {test_loss:.4f}')
# 保存权重
torch.save(
cvae.state_dict(),
PT_PATH
)
print(f"Saving PTH to:{PT_PATH}")
# 保存每轮损失
import pandas as pd
df = pd.DataFrame({
'Epoch': range(1, len(train_loss_list) + 1),
'Training Loss': train_loss_list,
'Test Loss': test_loss_list
})
df.to_csv(LOSS_PATH, index=False)
print(f"Saving Loss to:{LOSS_PATH}")训练详细信息:
Load Dataset......
==================================================
Train dataset size: 90000 samples
Test dataset size: 10000 samples
Train loader batch count: 90
Test loader batch count: 10
==================================================
Training the model...
Epoch [1/200], train loss 747.6686; val loss 213.2999
Epoch [2/200], train loss 139.1853; val loss 109.0857
Epoch [3/200], train loss 99.8494; val loss 93.2989
Epoch [4/200], train loss 89.7221; val loss 87.0099
......
Epoch [196/200], train loss 36.0690; val loss 36.1617
Epoch [197/200], train loss 35.9917; val loss 36.0512
Epoch [198/200], train loss 35.9444; val loss 36.0481
Epoch [199/200], train loss 35.8941; val loss 35.9624
Epoch [200/200], train loss 35.8199; val loss 35.9190
Saving PTH to:./result/model.pth
Saving Loss to:./result/losses.csv损失函数变化曲线: 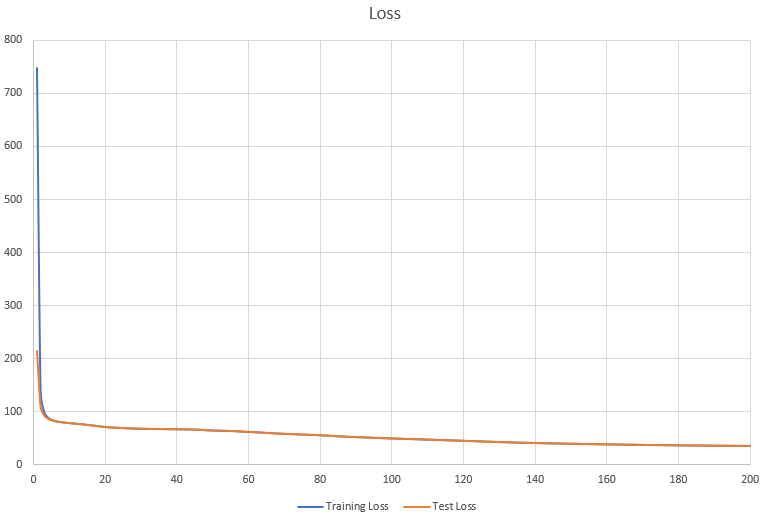
生成
以下代码在generate.py中实现,主要的差异在生成条件的独热编码,并输入模型(代码标黄部分)。
import numpy as np
import pandas as pd
from rdkit import Chem
from rdkit.Chem.Crippen import MolLogP, MolMR
import torch
from config import *
from model import CVAE
from rdkit import RDLogger
RDLogger.DisableLog('rdApp.*')
import warnings
warnings.filterwarnings(action='ignore')
# 分子特征矩阵转为mol对象
def graph_to_mol(node_labels, adjacency, atom_decoder_m, bond_decoder_m, strict=False):
mol = Chem.RWMol()
for node_label in node_labels:
mol.AddAtom(Chem.Atom(atom_decoder_m[node_label]))
for start, end in zip(*np.nonzero(adjacency)):
if start < end:
mol.AddBond(int(start), int(end), bond_decoder_m[adjacency[start, end]])
if strict:
try:
Chem.SanitizeMol(mol)
except:
mol = None
return mol
# 解析推理得到的数据
def cvae_results(
vae,
logP,
MR,
atom_decoder_m,
bond_decoder_m,
gen_num=GEN_NUM,
z_dim=Z_DIM,
cond_DIM=COND_DIM,
atom_types=ATOM_LEN,
bond_types=BOND_LEN,
row_dim=MAX_SIZE,
col_dim=F_COL,
):
with torch.no_grad():
z = torch.randn(int(gen_num), z_dim).to(DEVICE)
c1 = torch.zeros(int(gen_num), cond_DIM).to(DEVICE)
c1[:,logP] = 1
c2 = torch.zeros(int(gen_num), cond_DIM).to(DEVICE)
c2[:,MR] = 1
sample = vae.decoder(z, c1, c2)
smi = []
logp = []
mr = []
for matrix in sample.view(int(gen_num), 1, row_dim, col_dim).cpu():
try:
# 单个分子特征矩阵
matrix = matrix[0]
# 原子数编码矩阵
atom_num = matrix[:, 0]
# 独热转特征
atom_num = torch.argmax(atom_num).item() + 1
# 原子类型编码矩阵
atom_type = matrix[:,1:atom_types+1]
# 原子类型独热编码转类型索引
atom_type = torch.max(atom_type, -1)[1]
# 边特征编码矩阵
bonds = matrix[:, atom_types+1:]
# 边特征独热编码转特征索引的邻接矩阵
bonds = bonds.reshape(row_dim, row_dim, bond_types)
bonds = torch.max(bonds, dim=-1)[1]
# 只保留有效原子部分
nodes_hard_max = atom_type[:atom_num]
edges_hard_max = bonds[:atom_num, :atom_num]
# 从特征矩阵解码还原分子mol对象
mol = graph_to_mol(
nodes_hard_max.numpy(),
edges_hard_max.numpy(),
atom_decoder_m,
bond_decoder_m,
strict=True
)
# 有效性检查
if mol and '.' not in Chem.MolToSmiles(mol):
smi.append(Chem.MolToSmiles(mol))
logp.append(MolLogP(mol))
mr.append(MolMR(mol))
except Exception as e:
print(e)
continue
return pd.DataFrame({'SMILES': smi, 'logP': logp, 'MR': mr})最后把设定的条件输入模型,逐个组合生成需要的分子。
if __name__ == '__main__':
# 定义模型,迁移设备
cvae = CVAE()
cvae.to(DEVICE)
# 加载权重
cvae.load_state_dict(torch.load(PT_PATH, map_location=DEVICE), strict=True)
# 条件列
logP_list = CONDITIONS['log_list']
mr_list = CONDITIONS['mr_list']
conditions = [(logp, mr) for logp in logP_list for mr in mr_list]
print('Generating molecules...')
# 生成所有组合
for logp, mr in conditions:
cvae_df = cvae_results(cvae, int(logp), int(mr), atom_decoder_m, bond_decoder_m)
cvae_df.to_csv(f'{RESTULT_PATH}_{logp}_{mr}.csv', index=False)
print(f'Saving {RESTULT_PATH}_{logp}_{mr}.csv ({cvae_df.shape[0]})...')最终会生成多个给定条件的分子文件:
Generating molecules...
Saving ./result/result_0_2.csv (1531)...
Saving ./result/result_0_3.csv (3137)...
Saving ./result/result_0_4.csv (3533)...
Saving ./result/result_0_5.csv (3440)...数据分析
最后对模型生成的新分子性质分布做可视化查看,该部分代码在draw.py中实现:
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import matplotlib
from config import *
matplotlib.interactive(True)
# 数据加载
zinc = pd.read_csv('./data/ZINC.csv')
zinc = zinc[zinc['Length'] <= 16].reset_index(drop=True)
zinc_logP = zinc['logP'].tolist()
zinc_MR = [i/10 for i in zinc['MR'].tolist()]
# 样式配置
plt.rcParams["font.family"] = "Arial"
plt.rcParams["font.size"] = 8
plt.rcParams["axes.linewidth"] = 0.8
colors = {
"zinc_logP": "#FFA07A",
"zinc_MR": "#87CEFA",
"cvae_logP": "#CD5C5C",
"cvae_MR": "#4682B4"
}
# 动态获取参数列表
# 从 CONDITIONS 中动态读取,兼容任意长度
logP_list = CONDITIONS.get('logP_list', [])
MR_list = CONDITIONS.get('mr_list', [])
# 动态计算子图布局
n_rows = len(logP_list) # 行数
n_cols = len(MR_list) # 列数
total_plots = n_rows * n_cols # 总子图数
# 动态调整画布尺寸
# 基础尺寸:每列3英寸,每行2英寸,可根据需求调整
fig_width = 3 * n_cols if n_cols * 3 < 18 else 18 # 最大宽度限制18英寸
fig_height = 2 * n_rows if n_rows * 2 < 12 else 12 # 最大高度限制12英寸
# 生成动态子图
fig, axes = plt.subplots(
n_rows, n_cols,
figsize=(fig_width, fig_height),
gridspec_kw={"hspace": 0.4, "wspace": 0.3}
)
# 处理单行列的情况
if n_rows == 1 and n_cols == 1:
axes = np.array([[axes]]) # 转为2D数组
elif n_rows == 1:
axes = axes.reshape(1, -1)
elif n_cols == 1:
axes = axes.reshape(-1, 1)
# 展平用于循环
axes_flat = axes.flatten()
# 填充子图
for idx, ax in enumerate(axes_flat):
# 计算当前子图对应的 logP 和 MR 索引
row_idx = idx // n_cols
col_idx = idx % n_cols
target_logP = logP_list[row_idx]
target_MR = MR_list[col_idx]
try:
df = pd.read_csv(f'./result/result_{target_logP}_{target_MR}.csv')
cvae_logP = df['logP'].tolist()
cvae_MR = [i/10 for i in df['MR'].tolist()]
except FileNotFoundError:
print(f"警告:未找到文件 result_{target_logP}_{target_MR}.csv,跳过该子图")
ax.axis('off')
continue
# 绘制核密度曲线
sns.kdeplot(zinc_logP, color=colors["zinc_logP"], fill=True, alpha=0.5, ax=ax, label="_nolegend_")
sns.kdeplot(zinc_MR, color=colors["zinc_MR"], fill=True, alpha=0.5, ax=ax, label="_nolegend_")
sns.kdeplot(cvae_logP, color=colors["cvae_logP"], fill=True, alpha=0.5, ax=ax, label="_nolegend_")
sns.kdeplot(cvae_MR, color=colors["cvae_MR"], fill=True, alpha=0.5, ax=ax, label="_nolegend_")
# 绘制目标值垂直线
ax.axvline(x=-target_logP, color="red", linestyle="--", linewidth=1)
ax.axvline(x=target_MR, color="blue", linestyle="--", linewidth=1)
# 设置子图标题和轴属性
ax.set_title(f"logP={target_logP}, MR/10={target_MR}", fontsize=9)
ax.set_xlim(-5, 10)
ax.set_ylim(0, 1.4)
ax.set_xlabel("")
ax.set_ylabel("Density", fontsize=7)
ax.tick_params(axis="both", labelsize=7)
# 图例配置
handles = [
plt.Line2D([0], [0], color=colors["zinc_logP"], lw=2, label="ZINC (logP)"),
plt.Line2D([0], [0], color=colors["zinc_MR"], lw=2, label="ZINC (MR/10)"),
plt.Line2D([0], [0], color=colors["cvae_logP"], lw=2, label="CVAE (logP)"),
plt.Line2D([0], [0], color=colors["cvae_MR"], lw=2, label="CVAE (MR/10)")
]
fig.legend(
handles=handles,
labels=[h.get_label() for h in handles],
loc="center left",
bbox_to_anchor=(1.0, 0.5),
fontsize=9,
frameon=True,
borderaxespad=0.1
)
# 布局调整
plt.subplots_adjust(right=0.95)
plt.tight_layout() # 自动调整子图间距
plt.savefig('./result/generate.png', dpi=300, bbox_inches='tight')
plt.show(block=True)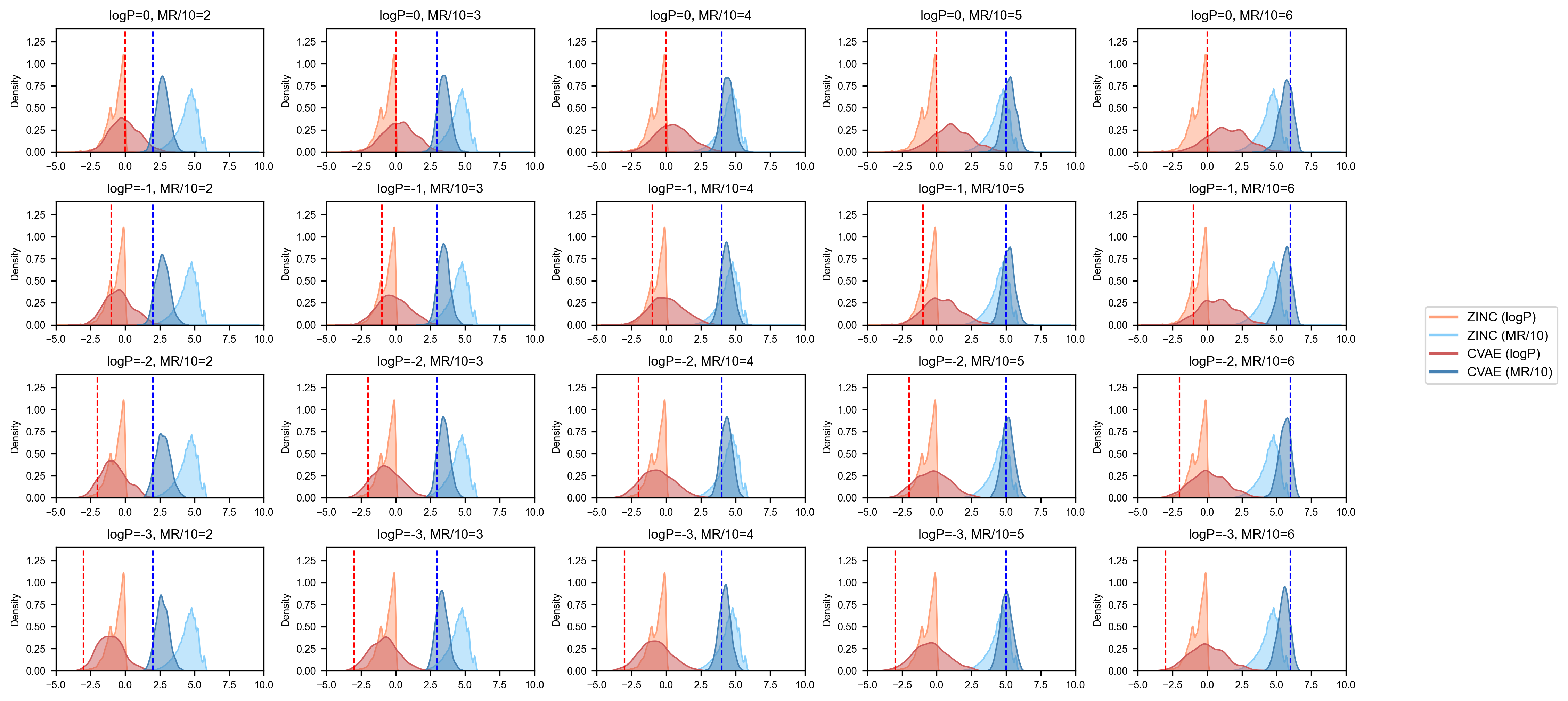
图中深色曲线是模型生成的数据分布,纵向标注线是我们指定的生成条件,可以看到对于MR条件,模型生成的分布和我们给定的条件基本一致,而logP分布匹配一般,这主要是因为训练数据集较少,后续可以尝试全量数据集训练,以达到最佳效果。